Funny SQL bug because I thought row_number() started at 0
I was working on a toy project of mine and ran into a funny SQL bug because of an off-by-one error.
Can you spot the error in this SQL docstring?
/* Aggregate features to arrays of size sequence_length (default 100) per user.
*
* userid, floor(row_number/100), feat1, feat2, feat3
* user1, 0 , [...], [...], [...]
* user1, 1 , [...], [...], [...]
* user2, 0 , [...], [...], [...]
*
* Define bucket_num = floor(row_number/100). This maps row 1-100 to 0, row
* 101-200 to 1, etc. Finally, ARRAY_AGG and GROUP BY the userid *and* the
* bucket_num.
* /
This docstring is part of my data prep routine that aggregates a row-based SQL dataset into feature arrays to use as input for my machine learning model.
First, I assign a number to each observation per user [0, 1, 2, ...]
select row_number() over (partition by userid) as row_number
Then, I create a variable that maps these numbers into buckets of 100 observations each
select floor(row_number/100) as bucket_num
Finally, I group the observations with the same userid and bucket number together and aggregate them into arrays
select array_agg(feature1) as feat1
...
group by userid, bucket_num
having count(*)=100
This is how I thought it was going to work:
| userid | row | row_number | floor(row_number/100) |
|---|---|---|---|
| 1 | 1 | 0 | 0 |
| 1 | 100 | 99 | 0 |
| 1 | 101 | 100 | 1 |
But when I started validating both datasets the aggregated dataset with feature arrays was missing 15% of users! What!? Weird.
So, what do you do here? I started out by trying some low-hanging fruit, mostly
stray where clauses. But none of them really seemed to hit the mark. However, once I started looking at the behavior of individual users everything started to make sense.
This is some data for a random user in the dataset that I’m using.
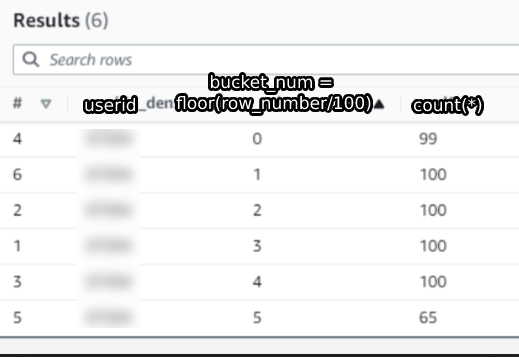
The first column is the userid, the second column is the floor(row_number/100) and the third column is a count of the rows in each bucket.
Surprisingly, we see that the first bucket only has 99 entries instead of the expected 100.
This anomaly showed immediately what was going wrong: Because the row number doesn’t start at 0 but starts at 1, we filter out users that have at max 2 sequences.
For example: Imagine a user with 150 rows in the dataset, because row number starts at 0 and not 1 we only map the first 99 rows to bucket 0 (1 to 99 is 99 numbers) and the remaining 51 rows to bucket 1. Both of these sequences are smaller than the required length of 100 and get filtered out by the having count(*)=100 clause.
We filter out both rows and this drops the whole user!
Thankfully, this is easy to fix. We simply subtract 1 from the row number to make the first bucket contain 100 elements:
| userid | row | row_number-1 | floor((row_number-1)/100) |
|---|---|---|---|
| 1 | 1 | 0 | 0 |
| 1 | .. | .. | .. |
| 1 | 100 | 99 | 0 |
| 1 | 101 | 100 | 1 |
Now we do not filter out users with only 2 sequences anymore because the first sequence, now with exactly 100 observatations, stays in the dataset. Problem solved!
Here is the corrected SQL snippet. The only difference is that we subtract 1 from the row number.
/* Aggregate features to arrays of size sequence_length (default 100) per user.
*
* userid, floor((row_number-1)/100), feat1, feat2, feat3
* user1, 0 , [...], [...], [...]
* user1, 1 , [...], [...], [...]
* user2, 0 , [...], [...], [...]
*
* Define bucket_num = floor((row_number-1)/100). This maps row 1-100 to 0, row
* 101-200 to 1, etc. Finally, ARRAY_AGG and GROUP BY the userid *and* the
* bucket_num.
* /
If you fiddle with row numbers, make sure to check whether they start at zero or one!




Comments